RNA_cormotif_Trajectory
2026-01-14
Last updated: 2026-01-28
Checks: 7 0
Knit directory: CrossSpecies_CM_Diff_RNA/
This reproducible R Markdown analysis was created with workflowr (version 1.7.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20251129) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 5a6d327. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: data/
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown
(analysis/RNA_Cormotif_Comp_Trajectory_Ensemble.Rmd) and
HTML (docs/RNA_Cormotif_Comp_Trajectory_Ensemble.html)
files. If you’ve configured a remote Git repository (see
?wflow_git_remote), click on the hyperlinks in the table
below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 5a6d327 | John D. Hurley | 2026-01-28 | Knit update |
| html | f388c90 | John D. Hurley | 2026-01-28 | Build site. |
| Rmd | 293a5c1 | John D. Hurley | 2026-01-28 | Updates for the website |
| Rmd | 085c1db | John D. Hurley | 2026-01-28 | Finalizing CorHeatMap |
knitr::opts_chunk$set(echo = TRUE,warning=FALSE,message=FALSE, dev=c("png","pdf"))####Library Loading####
library("edgeR")
library("ggplot2")
library("tibble")
library("dplyr")
library("ggrepel")
library("readr")
library("org.Hs.eg.db")
library("AnnotationDbi")
library("pheatmap")
library("tidyverse")
library("workflowr")
library("RUVSeq")
library("SummarizedExperiment")
library("readxl")
library("ggfortify")
library("ComplexHeatmap")
# BiocManager::install("SummarizedExperiment")
RNA_fc_df <- readRDS("data/Raw_Data/RNA_fc_df.RDS")
RNA_Metadata <- readRDS("data/Raw_Data/RNA_Metadata.RDS")
ann_colors <- readRDS("data/QC/ann_colors.RDS")
RNA_Metadata_No4 <- readRDS("data/QC/RNA_Metatdata_No4.RDS")
Filt_RMG0_RNA_fc_NoD4 <- readRDS("data/QC/Filt_RMG0_RNA_fc_NoD4.RDS")
Filt_RMG0_RNA_log2cpm_NoD4 <- readRDS("data/QC/Filt_RMG0_RNA_log2cpm_NoD4.RDS")
ann_colors_No4 <- readRDS("data/QC/ann_colors_no4.RDS")
Filt_RMG0_RNA_fc_NoD4_NoRep <- readRDS("data/Cormotif/Filt_RMG0_RNA_fc_NoD4_NoRep.RDS")
Filt_RMG0_RNA_log2cpm_NoD4_NoRep <- readRDS("data/Cormotif/Filt_RMG0_RNA_log2cpm_NoD4_NoRep.RDS")## Fit limma model using code as it is found in the original cormotif code. It has
## only been modified to add names to the matrix of t values, as well as the
## limma fits
limmafit.default <- function(exprs,groupid,compid) {
limmafits <- list()
compnum <- nrow(compid)
genenum <- nrow(exprs)
limmat <- matrix(0,genenum,compnum)
limmas2 <- rep(0,compnum)
limmadf <- rep(0,compnum)
limmav0 <- rep(0,compnum)
limmag1num <- rep(0,compnum)
limmag2num <- rep(0,compnum)
rownames(limmat) <- rownames(exprs)
colnames(limmat) <- rownames(compid)
names(limmas2) <- rownames(compid)
names(limmadf) <- rownames(compid)
names(limmav0) <- rownames(compid)
names(limmag1num) <- rownames(compid)
names(limmag2num) <- rownames(compid)
for(i in 1:compnum) {
selid1 <- which(groupid == compid[i,1])
selid2 <- which(groupid == compid[i,2])
eset <- new("ExpressionSet", exprs=cbind(exprs[,selid1],exprs[,selid2]))
g1num <- length(selid1)
g2num <- length(selid2)
designmat <- cbind(base=rep(1,(g1num+g2num)), delta=c(rep(0,g1num),rep(1,g2num)))
fit <- lmFit(eset,designmat)
fit <- eBayes(fit)
limmat[,i] <- fit$t[,2]
limmas2[i] <- fit$s2.prior
limmadf[i] <- fit$df.prior
limmav0[i] <- fit$var.prior[2]
limmag1num[i] <- g1num
limmag2num[i] <- g2num
limmafits[[i]] <- fit
# log odds
# w<-sqrt(1+fit$var.prior[2]/(1/g1num+1/g2num))
# log(0.99)+dt(fit$t[1,2],g1num+g2num-2+fit$df.prior,log=TRUE)-log(0.01)-dt(fit$t[1,2]/w, g1num+g2num-2+fit$df.prior, log=TRUE)+log(w)
}
names(limmafits) <- rownames(compid)
limmacompnum<-nrow(compid)
result<-list(t = limmat,
v0 = limmav0,
df0 = limmadf,
s20 = limmas2,
g1num = limmag1num,
g2num = limmag2num,
compnum = limmacompnum,
fits = limmafits)
}
limmafit.counts <-
function (exprs, groupid, compid, norm.factor.method = "TMM", voom.normalize.method = "none")
{
limmafits <- list()
compnum <- nrow(compid)
genenum <- nrow(exprs)
limmat <- matrix(NA,genenum,compnum)
limmas2 <- rep(0,compnum)
limmadf <- rep(0,compnum)
limmav0 <- rep(0,compnum)
limmag1num <- rep(0,compnum)
limmag2num <- rep(0,compnum)
rownames(limmat) <- rownames(exprs)
colnames(limmat) <- rownames(compid)
names(limmas2) <- rownames(compid)
names(limmadf) <- rownames(compid)
names(limmav0) <- rownames(compid)
names(limmag1num) <- rownames(compid)
names(limmag2num) <- rownames(compid)
for (i in 1:compnum) {
message(paste("Running limma for comparision",i,"/",compnum))
selid1 <- which(groupid == compid[i, 1])
selid2 <- which(groupid == compid[i, 2])
# make a new count data frame
counts <- cbind(exprs[, selid1], exprs[, selid2])
# remove NAs
not.nas <- which(apply(counts, 1, function(x) !any(is.na(x))) == TRUE)
# runn voom/limma
d <- DGEList(counts[not.nas,])
d <- calcNormFactors(d, method = norm.factor.method)
g1num <- length(selid1)
g2num <- length(selid2)
designmat <- cbind(base = rep(1, (g1num + g2num)), delta = c(rep(0,
g1num), rep(1, g2num)))
y <- voom(d, designmat, normalize.method = voom.normalize.method)
fit <- lmFit(y, designmat)
fit <- eBayes(fit)
limmafits[[i]] <- fit
limmat[not.nas, i] <- fit$t[, 2]
limmas2[i] <- fit$s2.prior
limmadf[i] <- fit$df.prior
limmav0[i] <- fit$var.prior[2]
limmag1num[i] <- g1num
limmag2num[i] <- g2num
}
limmacompnum <- nrow(compid)
names(limmafits) <- rownames(compid)
result <- list(t = limmat,
v0 = limmav0,
df0 = limmadf,
s20 = limmas2,
g1num = limmag1num,
g2num = limmag2num,
compnum = limmacompnum,
fits = limmafits)
}
limmafit.list <-
function (fitlist, cmp.idx=2)
{
compnum <- length(fitlist)
genes <- c()
for (i in 1:compnum) genes <- unique(c(genes, rownames(fitlist[[i]])))
genenum <- length(genes)
limmat <- matrix(NA,genenum,compnum)
limmas2 <- rep(0,compnum)
limmadf <- rep(0,compnum)
limmav0 <- rep(0,compnum)
limmag1num <- rep(0,compnum)
limmag2num <- rep(0,compnum)
rownames(limmat) <- genes
colnames(limmat) <- names(fitlist)
names(limmas2) <- names(fitlist)
names(limmadf) <- names(fitlist)
names(limmav0) <- names(fitlist)
names(limmag1num) <- names(fitlist)
names(limmag2num) <- names(fitlist)
for (i in 1:compnum) {
this.t <- fitlist[[i]]$t[,cmp.idx]
limmat[names(this.t),i] <- this.t
limmas2[i] <- fitlist[[i]]$s2.prior
limmadf[i] <- fitlist[[i]]$df.prior
limmav0[i] <- fitlist[[i]]$var.prior[cmp.idx]
limmag1num[i] <- sum(fitlist[[i]]$design[,cmp.idx]==0)
limmag2num[i] <- sum(fitlist[[i]]$design[,cmp.idx]==1)
}
limmacompnum <- compnum
result <- list(t = limmat,
v0 = limmav0,
df0 = limmadf,
s20 = limmas2,
g1num = limmag1num,
g2num = limmag2num,
compnum = limmacompnum,
fits = limmafits)
}
## Rank genes based on statistics
generank<-function(x) {
xcol<-ncol(x)
xrow<-nrow(x)
result<-matrix(0,xrow,xcol)
z<-(1:1:xrow)
for(i in 1:xcol) {
y<-sort(x[,i],decreasing=TRUE,na.last=TRUE)
result[,i]<-match(x[,i],y)
result[,i]<-order(result[,i])
}
result
}
## Log-likelihood for moderated t under H0
modt.f0.loglike<-function(x,df) {
a<-dt(x, df, log=TRUE)
result<-as.vector(a)
flag<-which(is.na(result)==TRUE)
result[flag]<-0
result
}
## Log-likelihood for moderated t under H1
## param=c(df,g1num,g2num,v0)
modt.f1.loglike<-function(x,param) {
df<-param[1]
g1num<-param[2]
g2num<-param[3]
v0<-param[4]
w<-sqrt(1+v0/(1/g1num+1/g2num))
dt(x/w, df, log=TRUE)-log(w)
a<-dt(x/w, df, log=TRUE)-log(w)
result<-as.vector(a)
flag<-which(is.na(result)==TRUE)
result[flag]<-0
result
}
## Correlation Motif Fit
cmfit.X<-function(x, type, K=1, tol=1e-3, max.iter=100) {
## initialize
xrow <- nrow(x)
xcol <- ncol(x)
loglike0 <- list()
loglike1 <- list()
p <- rep(1, K)/K
q <- matrix(runif(K * xcol), K, xcol)
q[1, ] <- rep(0.01, xcol)
for (i in 1:xcol) {
f0 <- type[[i]][[1]]
f0param <- type[[i]][[2]]
f1 <- type[[i]][[3]]
f1param <- type[[i]][[4]]
loglike0[[i]] <- f0(x[, i], f0param)
loglike1[[i]] <- f1(x[, i], f1param)
}
condlike <- list()
for (i in 1:xcol) {
condlike[[i]] <- matrix(0, xrow, K)
}
loglike.old <- -1e+10
for (i.iter in 1:max.iter) {
if ((i.iter%%50) == 0) {
print(paste("We have run the first ", i.iter, " iterations for K=",
K, sep = ""))
}
err <- tol + 1
clustlike <- matrix(0, xrow, K)
#templike <- matrix(0, xrow, 2)
templike1 <- rep(0, xrow)
templike2 <- rep(0, xrow)
for (j in 1:K) {
for (i in 1:xcol) {
templike1 <- log(q[j, i]) + loglike1[[i]]
templike2 <- log(1 - q[j, i]) + loglike0[[i]]
tempmax <- Rfast::Pmax(templike1, templike2)
templike1 <- exp(templike1 - tempmax)
templike2 <- exp(templike2 - tempmax)
tempsum <- templike1 + templike2
clustlike[, j] <- clustlike[, j] + tempmax +
log(tempsum)
condlike[[i]][, j] <- templike1/tempsum
}
clustlike[, j] <- clustlike[, j] + log(p[j])
}
#tempmax <- apply(clustlike, 1, max)
tempmax <- Rfast::rowMaxs(clustlike, value=TRUE)
for (j in 1:K) {
clustlike[, j] <- exp(clustlike[, j] - tempmax)
}
#tempsum <- apply(clustlike, 1, sum)
tempsum <- Rfast::rowsums(clustlike)
for (j in 1:K) {
clustlike[, j] <- clustlike[, j]/tempsum
}
#p.new <- (apply(clustlike, 2, sum) + 1)/(xrow + K)
p.new <- (Rfast::colsums(clustlike) + 1)/(xrow + K)
q.new <- matrix(0, K, xcol)
for (j in 1:K) {
clustpsum <- sum(clustlike[, j])
for (i in 1:xcol) {
q.new[j, i] <- (sum(clustlike[, j] * condlike[[i]][,
j]) + 1)/(clustpsum + 2)
}
}
err.p <- max(abs(p.new - p)/p)
err.q <- max(abs(q.new - q)/q)
err <- max(err.p, err.q)
loglike.new <- (sum(tempmax + log(tempsum)) + sum(log(p.new)) +
sum(log(q.new) + log(1 - q.new)))/xrow
p <- p.new
q <- q.new
loglike.old <- loglike.new
if (err < tol) {
break
}
}
clustlike <- matrix(0, xrow, K)
for (j in 1:K) {
for (i in 1:xcol) {
templike1 <- log(q[j, i]) + loglike1[[i]]
templike2 <- log(1 - q[j, i]) + loglike0[[i]]
tempmax <- Rfast::Pmax(templike1, templike2)
templike1 <- exp(templike1 - tempmax)
templike2 <- exp(templike2 - tempmax)
tempsum <- templike1 + templike2
clustlike[, j] <- clustlike[, j] + tempmax + log(tempsum)
condlike[[i]][, j] <- templike1/tempsum
}
clustlike[, j] <- clustlike[, j] + log(p[j])
}
#tempmax <- apply(clustlike, 1, max)
tempmax <- Rfast::rowMaxs(clustlike, value=TRUE)
for (j in 1:K) {
clustlike[, j] <- exp(clustlike[, j] - tempmax)
}
#tempsum <- apply(clustlike, 1, sum)
tempsum <- Rfast::rowsums(clustlike)
for (j in 1:K) {
clustlike[, j] <- clustlike[, j]/tempsum
}
p.post <- matrix(0, xrow, xcol)
for (j in 1:K) {
for (i in 1:xcol) {
p.post[, i] <- p.post[, i] + clustlike[, j] * condlike[[i]][,
j]
}
}
loglike.old <- loglike.old - (sum(log(p)) + sum(log(q) +
log(1 - q)))/xrow
loglike.old <- loglike.old * xrow
result <- list(p.post = p.post, motif.prior = p, motif.q = q,
loglike = loglike.old, clustlike=clustlike, condlike=condlike)
}
## Fit using (0,0,...,0) and (1,1,...,1)
cmfitall<-function(x, type, tol=1e-3, max.iter=100) {
## initialize
xrow<-nrow(x)
xcol<-ncol(x)
loglike0<-list()
loglike1<-list()
p<-0.01
## compute loglikelihood
L0<-matrix(0,xrow,1)
L1<-matrix(0,xrow,1)
for(i in 1:xcol) {
f0<-type[[i]][[1]]
f0param<-type[[i]][[2]]
f1<-type[[i]][[3]]
f1param<-type[[i]][[4]]
loglike0[[i]]<-f0(x[,i],f0param)
loglike1[[i]]<-f1(x[,i],f1param)
L0<-L0+loglike0[[i]]
L1<-L1+loglike1[[i]]
}
## EM algorithm to get MLE of p and q
loglike.old <- -1e10
for(i.iter in 1:max.iter) {
if((i.iter%%50) == 0) {
print(paste("We have run the first ", i.iter, " iterations",sep=""))
}
err<-tol+1
## compute posterior cluster membership
clustlike<-matrix(0,xrow,2)
clustlike[,1]<-log(1-p)+L0
clustlike[,2]<-log(p)+L1
tempmax<-apply(clustlike,1,max)
for(j in 1:2) {
clustlike[,j]<-exp(clustlike[,j]-tempmax)
}
tempsum<-apply(clustlike,1,sum)
## update motif occurrence rate
for(j in 1:2) {
clustlike[,j]<-clustlike[,j]/tempsum
}
p.new<-(sum(clustlike[,2])+1)/(xrow+2)
## evaluate convergence
err<-abs(p.new-p)/p
## evaluate whether the log.likelihood increases
loglike.new<-(sum(tempmax+log(tempsum))+log(p.new)+log(1-p.new))/xrow
loglike.old<-loglike.new
p<-p.new
if(err<tol) {
break;
}
}
## compute posterior p
clustlike<-matrix(0,xrow,2)
clustlike[,1]<-log(1-p)+L0
clustlike[,2]<-log(p)+L1
tempmax<-apply(clustlike,1,max)
for(j in 1:2) {
clustlike[,j]<-exp(clustlike[,j]-tempmax)
}
tempsum<-apply(clustlike,1,sum)
for(j in 1:2) {
clustlike[,j]<-clustlike[,j]/tempsum
}
p.post<-matrix(0,xrow,xcol)
for(i in 1:xcol) {
p.post[,i]<-clustlike[,2]
}
## return
#calculate back loglikelihood
loglike.old<-loglike.old-(log(p)+log(1-p))/xrow
loglike.old<-loglike.old*xrow
result<-list(p.post=p.post, motif.prior=p, loglike=loglike.old)
}
## Fit each dataset separately
cmfitsep<-function(x, type, tol=1e-3, max.iter=100) {
## initialize
xrow<-nrow(x)
xcol<-ncol(x)
loglike0<-list()
loglike1<-list()
p<-0.01*rep(1,xcol)
loglike.final<-rep(0,xcol)
## compute loglikelihood
for(i in 1:xcol) {
f0<-type[[i]][[1]]
f0param<-type[[i]][[2]]
f1<-type[[i]][[3]]
f1param<-type[[i]][[4]]
loglike0[[i]]<-f0(x[,i],f0param)
loglike1[[i]]<-f1(x[,i],f1param)
}
p.post<-matrix(0,xrow,xcol)
## EM algorithm to get MLE of p
for(coli in 1:xcol) {
loglike.old <- -1e10
for(i.iter in 1:max.iter) {
if((i.iter%%50) == 0) {
print(paste("We have run the first ", i.iter, " iterations",sep=""))
}
err<-tol+1
## compute posterior cluster membership
clustlike<-matrix(0,xrow,2)
clustlike[,1]<-log(1-p[coli])+loglike0[[coli]]
clustlike[,2]<-log(p[coli])+loglike1[[coli]]
tempmax<-apply(clustlike,1,max)
for(j in 1:2) {
clustlike[,j]<-exp(clustlike[,j]-tempmax)
}
tempsum<-apply(clustlike,1,sum)
## evaluate whether the log.likelihood increases
loglike.new<-sum(tempmax+log(tempsum))/xrow
## update motif occurrence rate
for(j in 1:2) {
clustlike[,j]<-clustlike[,j]/tempsum
}
p.new<-(sum(clustlike[,2]))/(xrow)
## evaluate convergence
err<-abs(p.new-p[coli])/p[coli]
loglike.old<-loglike.new
p[coli]<-p.new
if(err<tol) {
break;
}
}
## compute posterior p
clustlike<-matrix(0,xrow,2)
clustlike[,1]<-log(1-p[coli])+loglike0[[coli]]
clustlike[,2]<-log(p[coli])+loglike1[[coli]]
tempmax<-apply(clustlike,1,max)
for(j in 1:2) {
clustlike[,j]<-exp(clustlike[,j]-tempmax)
}
tempsum<-apply(clustlike,1,sum)
for(j in 1:2) {
clustlike[,j]<-clustlike[,j]/tempsum
}
p.post[,coli]<-clustlike[,2]
loglike.final[coli]<-loglike.old
}
## return
loglike.final<-loglike.final*xrow
result<-list(p.post=p.post, motif.prior=p, loglike=loglike.final)
}
## Fit the full model
cmfitfull<-function(x, type, tol=1e-3, max.iter=100) {
## initialize
xrow<-nrow(x)
xcol<-ncol(x)
loglike0<-list()
loglike1<-list()
K<-2^xcol
p<-rep(1,K)/K
pattern<-rep(0,xcol)
patid<-matrix(0,K,xcol)
## compute loglikelihood
for(i in 1:xcol) {
f0<-type[[i]][[1]]
f0param<-type[[i]][[2]]
f1<-type[[i]][[3]]
f1param<-type[[i]][[4]]
loglike0[[i]]<-f0(x[,i],f0param)
loglike1[[i]]<-f1(x[,i],f1param)
}
L<-matrix(0,xrow,K)
for(i in 1:K)
{
patid[i,]<-pattern
for(j in 1:xcol) {
if(pattern[j] < 0.5) {
L[,i]<-L[,i]+loglike0[[j]]
} else {
L[,i]<-L[,i]+loglike1[[j]]
}
}
if(i < K) {
pattern[xcol]<-pattern[xcol]+1
j<-xcol
while(pattern[j] > 1) {
pattern[j]<-0
j<-j-1
pattern[j]<-pattern[j]+1
}
}
}
## EM algorithm to get MLE of p and q
loglike.old <- -1e10
for(i.iter in 1:max.iter) {
if((i.iter%%50) == 0) {
print(paste("We have run the first ", i.iter, " iterations",sep=""))
}
err<-tol+1
## compute posterior cluster membership
clustlike<-matrix(0,xrow,K)
for(j in 1:K) {
clustlike[,j]<-log(p[j])+L[,j]
}
tempmax<-apply(clustlike,1,max)
for(j in 1:K) {
clustlike[,j]<-exp(clustlike[,j]-tempmax)
}
tempsum<-apply(clustlike,1,sum)
## update motif occurrence rate
for(j in 1:K) {
clustlike[,j]<-clustlike[,j]/tempsum
}
p.new<-(apply(clustlike,2,sum)+1)/(xrow+K)
## evaluate convergence
err<-max(abs(p.new-p)/p)
## evaluate whether the log.likelihood increases
loglike.new<-(sum(tempmax+log(tempsum))+sum(log(p.new)))/xrow
loglike.old<-loglike.new
p<-p.new
if(err<tol) {
break;
}
}
## compute posterior p
clustlike<-matrix(0,xrow,K)
for(j in 1:K) {
clustlike[,j]<-log(p[j])+L[,j]
}
tempmax<-apply(clustlike,1,max)
for(j in 1:K) {
clustlike[,j]<-exp(clustlike[,j]-tempmax)
}
tempsum<-apply(clustlike,1,sum)
for(j in 1:K) {
clustlike[,j]<-clustlike[,j]/tempsum
}
p.post<-matrix(0,xrow,xcol)
for(j in 1:K) {
for(i in 1:xcol) {
if(patid[j,i] > 0.5) {
p.post[,i]<-p.post[,i]+clustlike[,j]
}
}
}
## return
#calculate back loglikelihood
loglike.old<-loglike.old-sum(log(p))/xrow
loglike.old<-loglike.old*xrow
result<-list(p.post=p.post, motif.prior=p, loglike=loglike.old)
}
generatetype<-function(limfitted)
{
jtype<-list()
df<-limfitted$g1num+limfitted$g2num-2+limfitted$df0
for(j in 1:limfitted$compnum)
{
jtype[[j]]<-list(f0=modt.f0.loglike, f0.param=df[j], f1=modt.f1.loglike, f1.param=c(df[j],limfitted$g1num[j],limfitted$g2num[j],limfitted$v0[j]))
}
jtype
}
cormotiffit <- function(exprs, groupid=NULL, compid=NULL, K=1, tol=1e-3,
max.iter=100, BIC=TRUE, norm.factor.method="TMM",
voom.normalize.method = "none", runtype=c("logCPM","counts","limmafits"), each=3)
{
# first I want to do some typechecking. Input can be either a normalized
# matrix, a count matrix, or a list of limma fits. Dispatch the correct
# limmafit accordingly.
# todo: add some typechecking here
limfitted <- list()
if (runtype=="counts") {DOX_24T_shared_DEGs
limfitted <- limmafit.counts(exprs,groupid,compid, norm.factor.method, voom.normalize.method)
} else if (runtype=="logCPM") {
limfitted <- limmafit.default(exprs,groupid,compid)
} else if (runtype=="limmafits") {
limfitted <- limmafit.list(exprs)
} else {
stop("runtype must be one of 'logCPM', 'counts', or 'limmafits'")
}
jtype<-generatetype(limfitted)
fitresult<-list()
ks <- rep(K, each = each)
fitresult <- bplapply(1:length(ks), function(i, x, type, ks, tol, max.iter) {
cmfit.X(x, type, K = ks[i], tol = tol, max.iter = max.iter)
}, x=limfitted$t, type=jtype, ks=ks, tol=tol, max.iter=max.iter)
best.fitresults <- list()
for (i in 1:length(K)) {
w.k <- which(ks==K[i])
this.bic <- c()
for (j in w.k) this.bic[j] <- -2 * fitresult[[j]]$loglike + (K[i] - 1 + K[i] * limfitted$compnum) * log(dim(limfitted$t)[1])
w.min <- which(this.bic == min(this.bic, na.rm = TRUE))[1]
best.fitresults[[i]] <- fitresult[[w.min]]
}
fitresult <- best.fitresults
bic <- rep(0, length(K))
aic <- rep(0, length(K))
loglike <- rep(0, length(K))
for (i in 1:length(K)) loglike[i] <- fitresult[[i]]$loglike
for (i in 1:length(K)) bic[i] <- -2 * fitresult[[i]]$loglike + (K[i] - 1 + K[i] * limfitted$compnum) * log(dim(limfitted$t)[1])
for (i in 1:length(K)) aic[i] <- -2 * fitresult[[i]]$loglike + 2 * (K[i] - 1 + K[i] * limfitted$compnum)
if(BIC==TRUE) {
bestflag=which(bic==min(bic))
}
else {
bestflag=which(aic==min(aic))
}
result<-list(bestmotif=fitresult[[bestflag]],bic=cbind(K,bic),
aic=cbind(K,aic),loglike=cbind(K,loglike), allmotifs=fitresult)
}
cormotiffitall<-function(exprs,groupid,compid, tol=1e-3, max.iter=100)
{
limfitted<-limmafit(exprs,groupid,compid)
jtype<-generatetype(limfitted)
fitresult<-cmfitall(limfitted$t,type=jtype,tol=1e-3,max.iter=max.iter)
}
cormotiffitsep<-function(exprs,groupid,compid, tol=1e-3, max.iter=100)
{
limfitted<-limmafit(exprs,groupid,compid)
jtype<-generatetype(limfitted)
fitresult<-cmfitsep(limfitted$t,type=jtype,tol=1e-3,max.iter=max.iter)
}
cormotiffitfull<-function(exprs,groupid,compid, tol=1e-3, max.iter=100)
{
limfitted<-limmafit(exprs,groupid,compid)
jtype<-generatetype(limfitted)
fitresult<-cmfitfull(limfitted$t,type=jtype,tol=1e-3,max.iter=max.iter)
}
plotIC<-function(fitted_cormotif)
{
oldpar<-par(mfrow=c(1,2))
plot(fitted_cormotif$bic[,1], fitted_cormotif$bic[,2], type="b",xlab="Motif Number", ylab="BIC", main="BIC")
plot(fitted_cormotif$aic[,1], fitted_cormotif$aic[,2], type="b",xlab="Motif Number", ylab="AIC", main="AIC")
}
plotMotif<-function(fitted_cormotif,title="")
{
layout(matrix(1:2,ncol=2))
u<-1:dim(fitted_cormotif$bestmotif$motif.q)[2]
v<-1:dim(fitted_cormotif$bestmotif$motif.q)[1]
image(u,v,t(fitted_cormotif$bestmotif$motif.q),
col=gray(seq(from=1,to=0,by=-0.1)),xlab="Study",yaxt = "n",
ylab="Corr. Motifs",main=paste(title,"pattern",sep=" "))
axis(2,at=1:length(v))
for(i in 1:(length(u)+1))
{
abline(v=(i-0.5))
}
for(i in 1:(length(v)+1))
{
abline(h=(i-0.5))
}
Ng=10000
if(is.null(fitted_cormotif$bestmotif$p.post)!=TRUE)
Ng=nrow(fitted_cormotif$bestmotif$p.post)
genecount=floor(fitted_cormotif$bestmotif$motif.p*Ng)
NK=nrow(fitted_cormotif$bestmotif$motif.q)
plot(0,0.7,pch=".",xlim=c(0,1.2),ylim=c(0.75,NK+0.25),
frame.plot=FALSE,axes=FALSE,xlab="No. of genes",ylab="", main=paste(title,"frequency",sep=" "))
segments(0,0.7,fitted_cormotif$bestmotif$motif.p[1],0.7)
rect(0,1:NK-0.3,fitted_cormotif$bestmotif$motif.p,1:NK+0.3,
col="dark grey")
mtext(1:NK,at=1:NK,side=2,cex=0.8)
text(fitted_cormotif$bestmotif$motif.p+0.15,1:NK,
labels=floor(fitted_cormotif$bestmotif$motif.p*Ng))
}check_overlap <- function(list_of_lists) {
# create empty list to store results
overlap_results <- list()
# loop over each cluster
for (i in seq_along(list_of_lists)) {
cluster_name <- names(list_of_lists)[i]
genes_i <- list_of_lists[[i]]
# compare to all other clusters
overlaps <- sapply(seq_along(list_of_lists), function(j) {
if (i == j) return(NA) # skip self
genes_j <- list_of_lists[[j]]
length(intersect(genes_i, genes_j)) # count of overlapping genes
})
names(overlaps) <- names(list_of_lists)
overlap_results[[cluster_name]] <- overlaps
}
return(overlap_results)
}# Seed0 <- readRDS("~/diff_timeline_tes/RNA/Run1_Run2_Concat/cormotif/CorMotif_k1_k20_DayBefore_seed0.rds")
# Seed1 <- readRDS("~/diff_timeline_tes/RNA/Run1_Run2_Concat/cormotif/CorMotif_k1_k20_seed1.rds")
# Seed2 <- readRDS("~/diff_timeline_tes/RNA/Run1_Run2_Concat/cormotif/CorMotif_k1_k20_seed2.rds")
# Seed3 <- readRDS("~/diff_timeline_tes/RNA/Run1_Run2_Concat/cormotif/CorMotif_k1_k20_seed3.rds")
# Seed4 <- readRDS("~/diff_timeline_tes/RNA/Run1_Run2_Concat/cormotif/CorMotif_k1_k20_seed4.rds")
# Seed5 <- readRDS("~/diff_timeline_tes/RNA/Run1_Run2_Concat/cormotif/CorMotif_k1_k20_seed5.rds")
# Seed6 <- readRDS("~/diff_timeline_tes/RNA/Run1_Run2_Concat/cormotif/CorMotif_k1_k20_seed6.rds")
# Seed7 <- readRDS("~/diff_timeline_tes/RNA/Run1_Run2_Concat/cormotif/CorMotif_k1_k20_seed7.rds")
# Seed8 <- readRDS("~/diff_timeline_tes/RNA/Run1_Run2_Concat/cormotif/CorMotif_k1_k20_seed8.rds")
# Seed9 <- readRDS("~/diff_timeline_tes/RNA/Run1_Run2_Concat/cormotif/CorMotif_k1_k20_seed9.rds")
# Seed10 <- readRDS("~/diff_timeline_tes/RNA/Run1_Run2_Concat/cormotif/CorMotif_k1_k20_seed10.rds")
#
# CoMortiffits_TACC <- list(
# Seed0,
# Seed1,
# Seed2,
# Seed3,
# Seed4,
# Seed5,
# Seed6,
# Seed7,
# Seed8,
# Seed9,
# Seed10
# )
#
# #saveRDS(CoMortiffits_TACC,"data/Cormotif/DayBefore_CoMortiffits_TACC.RDS")
# Seed8$bic# extract_bic <- function(fit, seed_id) {
# # bic is a 20x2 matrix: column "bic" contains the actual BIC
# bic_values <- fit$bic[, "bic"]
#
# tibble(
# seed = seed_id,
# motifs = seq_along(bic_values),
# BIC = as.numeric(bic_values)
# )
# }
#
# # Define %||% if not using rlang
# `%||%` <- function(x, y) if(!is.null(x)) x else y
#
#
# names(DayBefore_CoMortiffits_TACC) <- paste0("seed_", seq_along(DayBefore_CoMortiffits_TACC))
#
# bic_table <- map2_dfr(
# DayBefore_CoMortiffits_TACC,
# names(DayBefore_CoMortiffits_TACC),
# extract_bic
# )
#
# head(bic_table)
# table(bic_table$seed)
#
# bic_summary <- bic_table %>%
# group_by(motifs) %>%
# summarise(
# mean_BIC = mean(BIC),
# sd_BIC = sd(BIC),
# n_seeds = n(),
# .groups = "drop"
# )
#
# bic_summary
#
# plotIC(Seed8)Filt_RMG0_RNA_log2cpm_NoD4_NoRep <- Filt_RMG0_RNA_log2cpm_NoD4 %>%
as.data.frame() %>% # convert to data frame
dplyr::select(-contains("R")) %>% # remove columns containing "R"
as.matrix() # convert back to matrix
# saveRDS(Filt_RMG0_RNA_log2cpm_NoD4_NoRep,"data/Cormotif/Filt_RMG0_RNA_log2cpm_NoD4_NoRep.RDS")groupid <- c(rep(1:5,7),rep(6:10,7))
compid_Cond1 <- c(1,2,3,4,6,7,8,9)
compid_Cond2 <- c(2,3,4,5,7,8,9,10)
compid <- cbind(compid_Cond1,compid_Cond2)
#Model Fitting
# set.seed(8)
# motif.fitted<-cormotiffit(exprs = Filt_RMG0_RNA_log2cpm_NoD4_NoRep,
# groupid = groupid,
# compid = compid,
# K=11,
# max.iter = 3000,
# runtype = "logCPM")
#
#
# saveRDS(motif.fitted,"data/Cormotif/cormotif_RNA__seed8_k8_DayBefore_emmaCode.RDS")
motif.fitted <- readRDS("data/Cormotif/cormotif_RNA__seed8_k8_DayBefore_emmaCode.RDS")
plotIC(motif.fitted)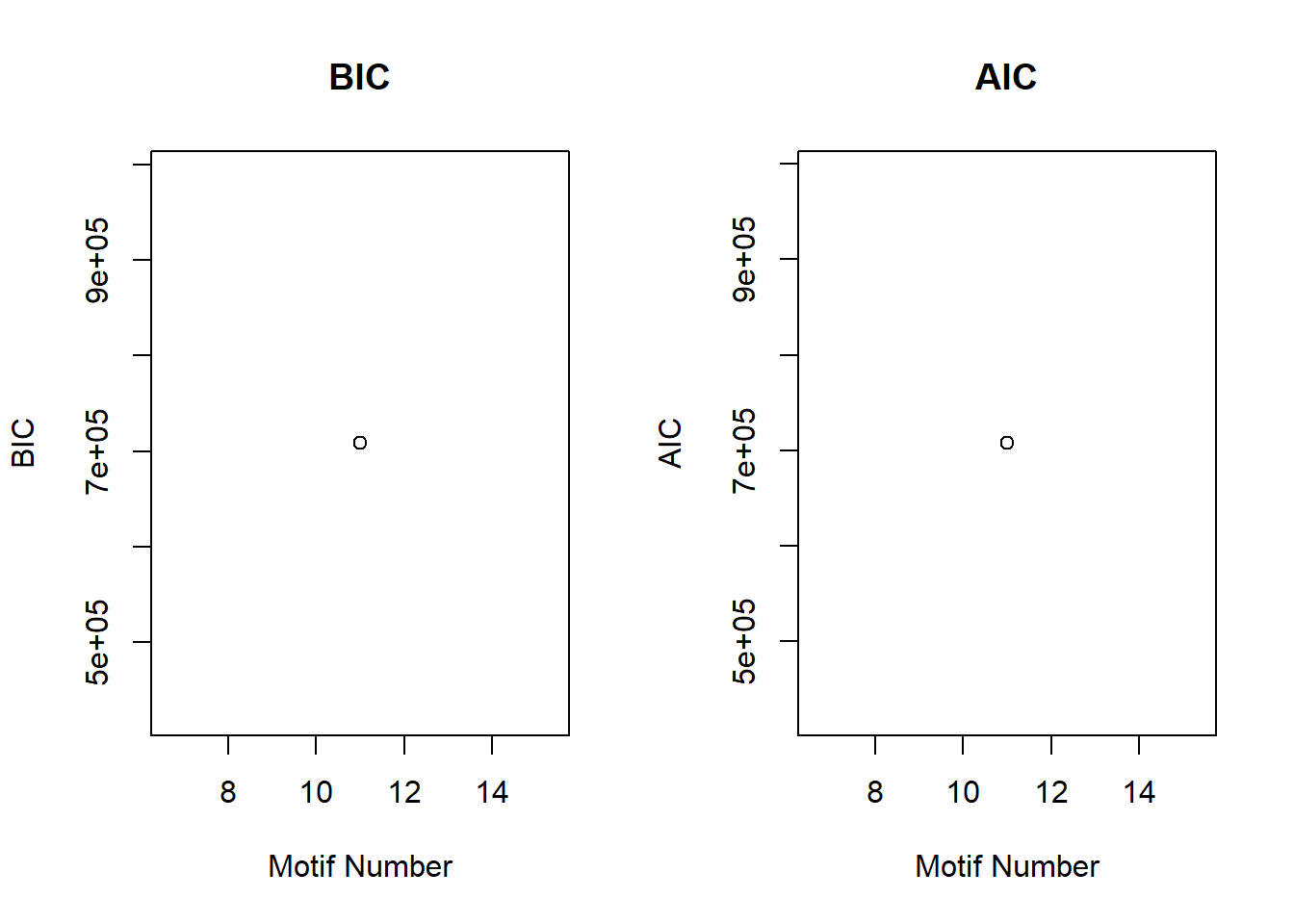
| Version | Author | Date |
|---|---|---|
| f388c90 | John D. Hurley | 2026-01-28 |
motif.fitted$bic K bic
[1,] 11 708372.2plotMotif(motif.fitted)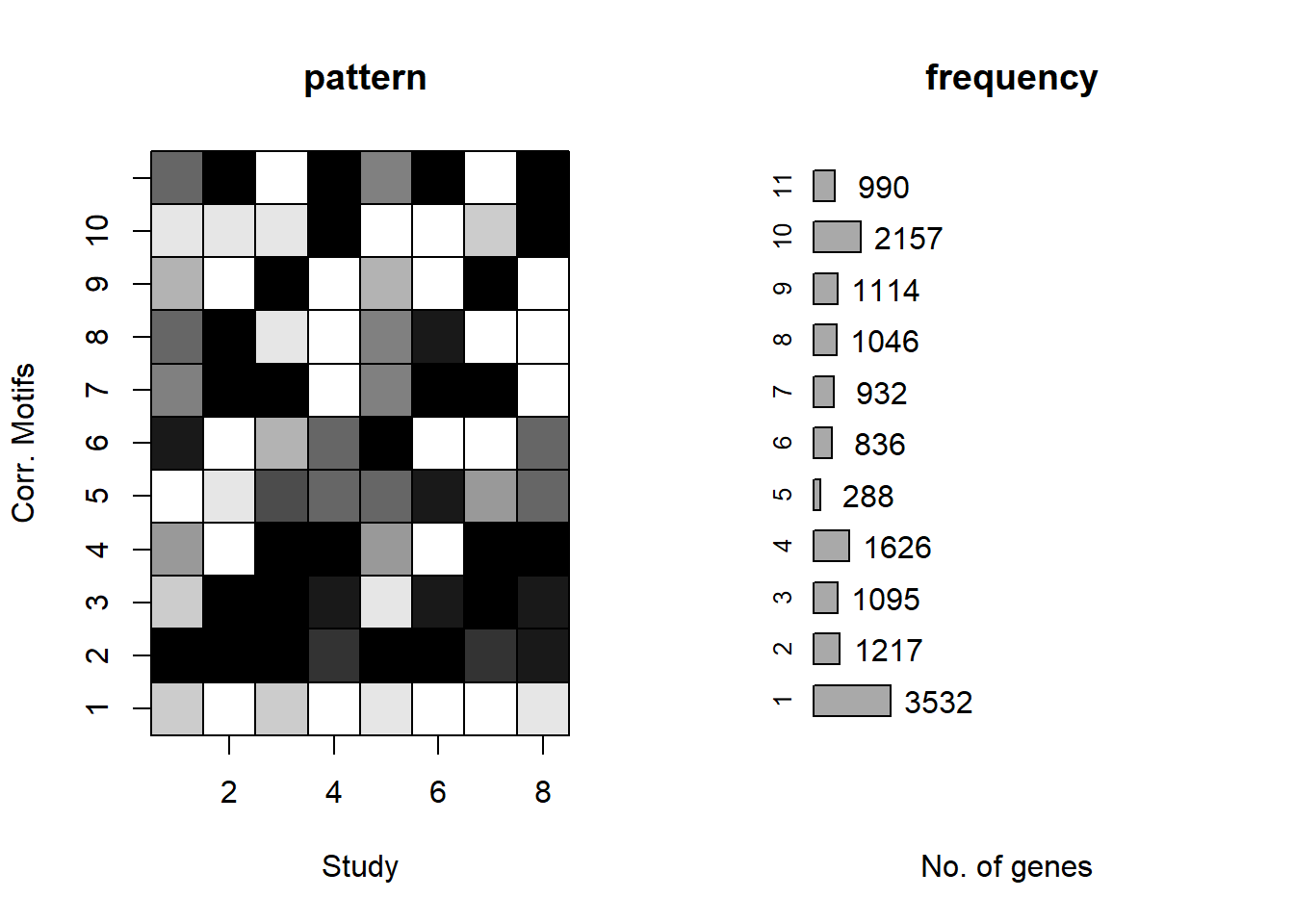
| Version | Author | Date |
|---|---|---|
| f388c90 | John D. Hurley | 2026-01-28 |
# Studies Human Day0 - 1. HDay2, 2. HDay5, 3. HDay15, 4. HDay30
# Chimp Day0 - 5. CDay2, 6. CDay5, 7. CDay15, 8. CDay30# Set up Variables
groupid <- c(rep(1:5,7),rep(6:10,7))
compid_Cond1 <- c(1,2,3,4,6,7,8,9)
compid_Cond2 <- c(2,3,4,5,7,8,9,10)
# Extract motif matrix
motif_matrix <- as.matrix(motif.fitted$bestmotif$motif.q)
# Label rows and columns
rownames(motif_matrix) <- paste0("Motif_", seq_len(nrow(motif_matrix)))
colnames(motif_matrix) <- paste0("C", compid[,1], "_vs_C", compid[,2])
# Convert to tidy format
motif_df <- as.data.frame(motif_matrix) %>%
mutate(Motif = rownames(.)) %>%
pivot_longer(-Motif, names_to = "Comparison", values_to = "Probability")
# Reorder motifs 1–11
motif_df$Motif <- factor(motif_df$Motif, levels = paste0("Motif_", 1:11))
motif_df$Comparison <- factor(motif_df$Comparison, levels = c("C1_vs_C2","C2_vs_C3","C3_vs_C4","C4_vs_C5","C6_vs_C7","C7_vs_C8","C8_vs_C9","C9_vs_C10"))
# Plot
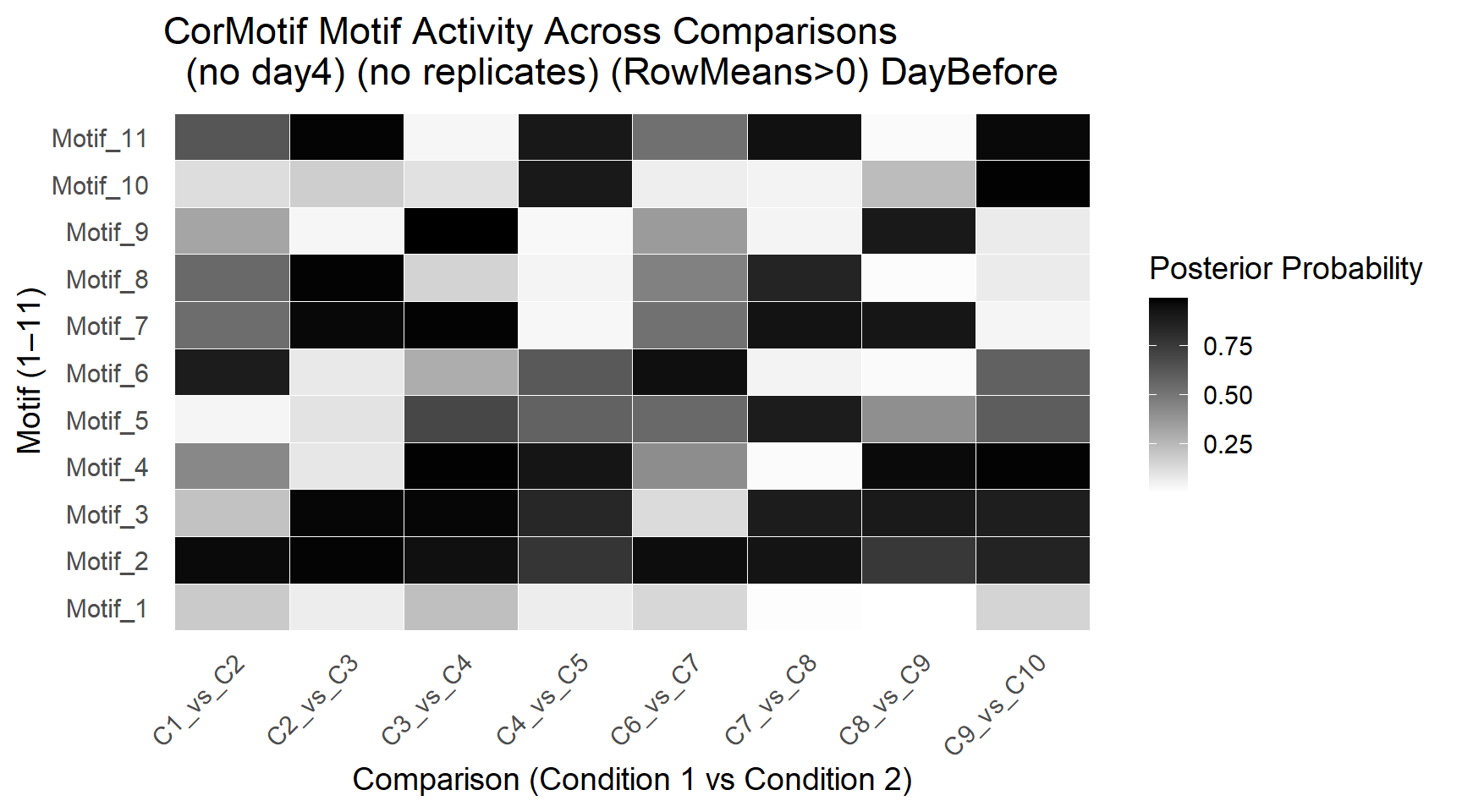
ggplot(motif_df, aes(x = Comparison, y = Motif, fill = Probability)) +
geom_tile(color = "white") +
scale_fill_gradient(low = "white", high = "black") +
labs(
title = "CorMotif Motif Activity Across Comparisons \n (no day4) (no replicates) (RowMeans>0) DayBefore" ,
x = "Comparison (Condition 1 vs Condition 2)",
y = "Motif (1–11)",
fill = "Posterior Probability"
) +
theme_minimal(base_size = 14) +
theme(
axis.text.x = element_text(angle = 45, hjust = 1),
panel.grid = element_blank()
)
| Version | Author | Date |
|---|---|---|
| f388c90 | John D. Hurley | 2026-01-28 |
# Comparison C1 = Human Day0
# Comparison C2 = Human Day2
# Comparison C3 = Human Day5
# Comparison C4 = Human Day15
# Comparison C5 = Human Day30
# Comparison C6 = Chimp Day0
# Comparison C7 = Chimp Day2
# Comparison C8 = Chimp Day5
# Comparison C9 = Chimp Day15
# Comparison C10 = Chimp Day30motif.fit <- motif.fitted
log2cpm <- Filt_RMG0_RNA_log2cpm_NoD4
#extract the posterior probability that these genes belong to motifs
gene_prob_ppost <- motif.fit$bestmotif$p.post
rownames(gene_prob_ppost) <- rownames(log2cpm)
# saveRDS(gene_prob_ppost,"data/Cormotif/RNA_gene_prob_ppost_Trajectory.RDS")
#assign each gene to a motif with max post prob
assigned_motifs <- apply(gene_prob_ppost, 1, which.max)
max_probs <- apply(gene_prob_ppost, 1, max)
#combine these into a dataframe - motif assigned genes (p.post)
motif_assignment_df <- gene_prob_ppost %>%
as.data.frame() %>%
rownames_to_column("Gene") %>%
mutate(
Assigned_Motif = assigned_motifs[Gene],
Max_Probability = max_probs[Gene]
)
#make some histograms of the unfiltered data from Cormotif p.post
gene_prob_ppost %>%
as.data.frame() %>%
ggplot(., aes(x = V1))+
geom_histogram(bins = 50)+
xlim(0,1)+
ggtitle("Cormtif p.post RNA V1")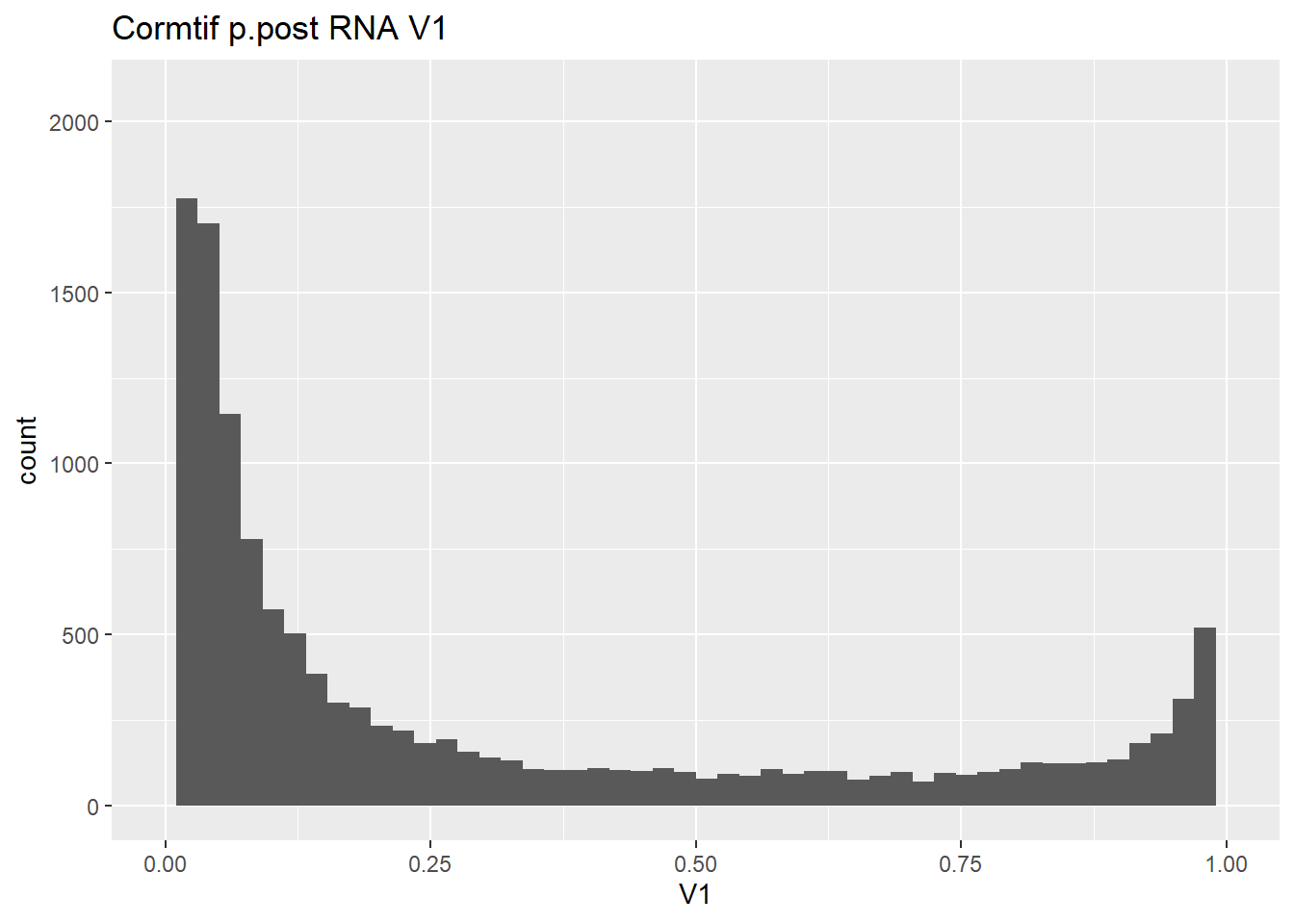
| Version | Author | Date |
|---|---|---|
| f388c90 | John D. Hurley | 2026-01-28 |
gene_prob_ppost %>%
as.data.frame() %>%
ggplot(., aes(x = V2))+
geom_histogram(bins = 50)+
xlim(0,1)+
ggtitle("Cormtif p.post RNA V2")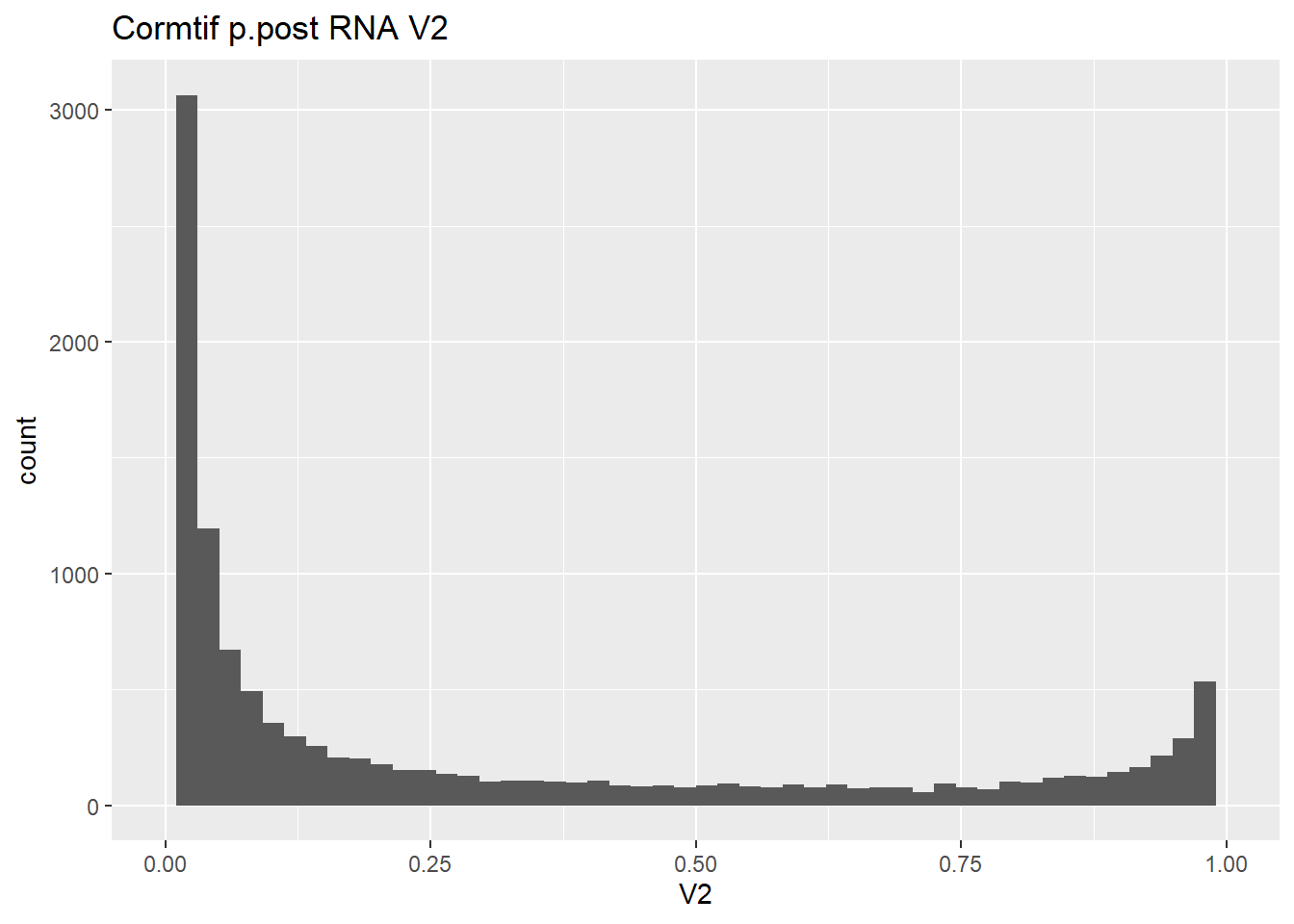
| Version | Author | Date |
|---|---|---|
| f388c90 | John D. Hurley | 2026-01-28 |
gene_prob_ppost %>%
as.data.frame() %>%
ggplot(., aes(x = V3))+
geom_histogram(bins = 50)+
xlim(0,1)+
ggtitle("Cormtif p.post RNA V3")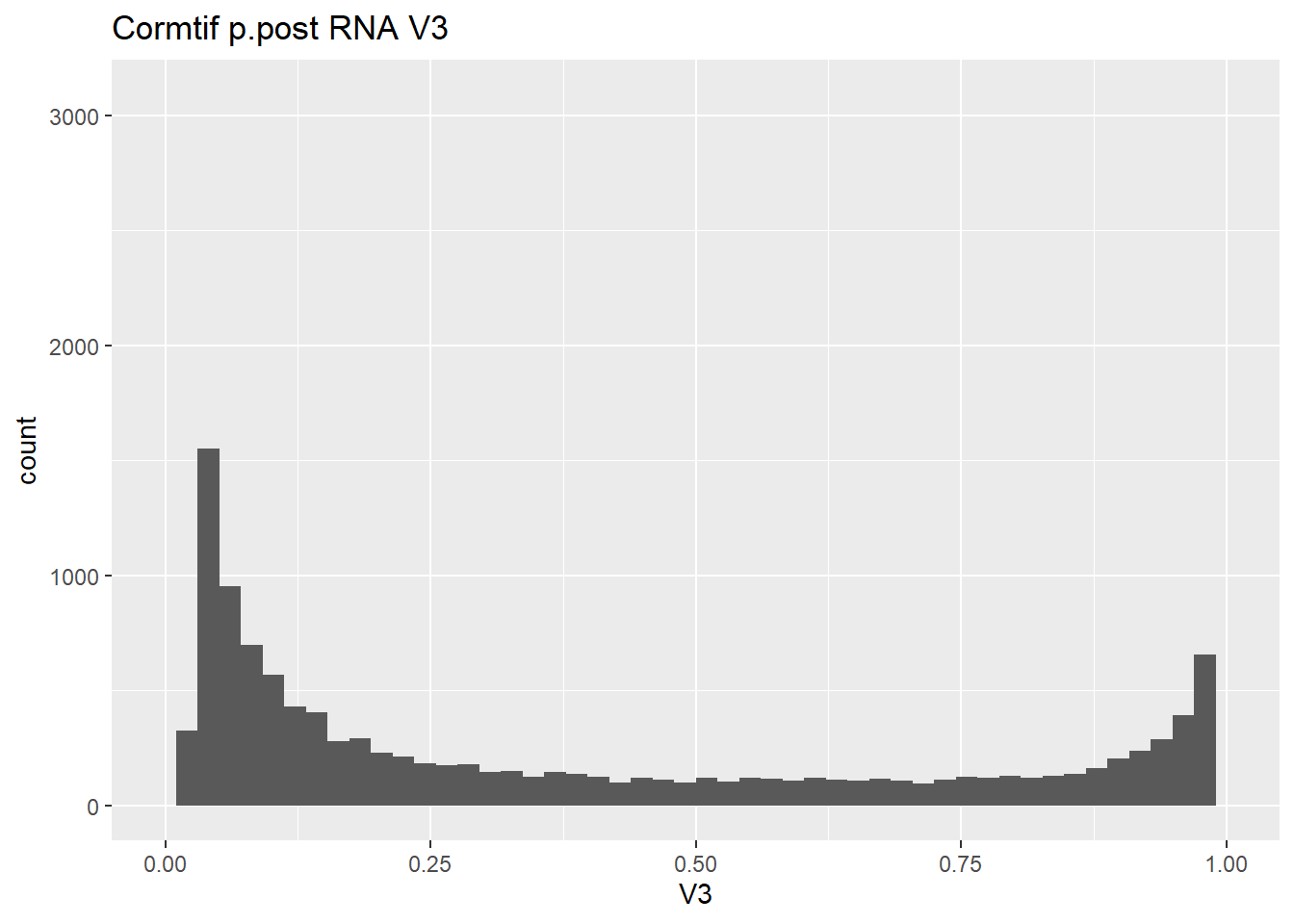
| Version | Author | Date |
|---|---|---|
| f388c90 | John D. Hurley | 2026-01-28 |
gene_prob_ppost %>%
as.data.frame() %>%
ggplot(., aes(x = V4))+
geom_histogram(bins = 50)+
xlim(0,1)+
ggtitle("Cormtif p.post RNA V4")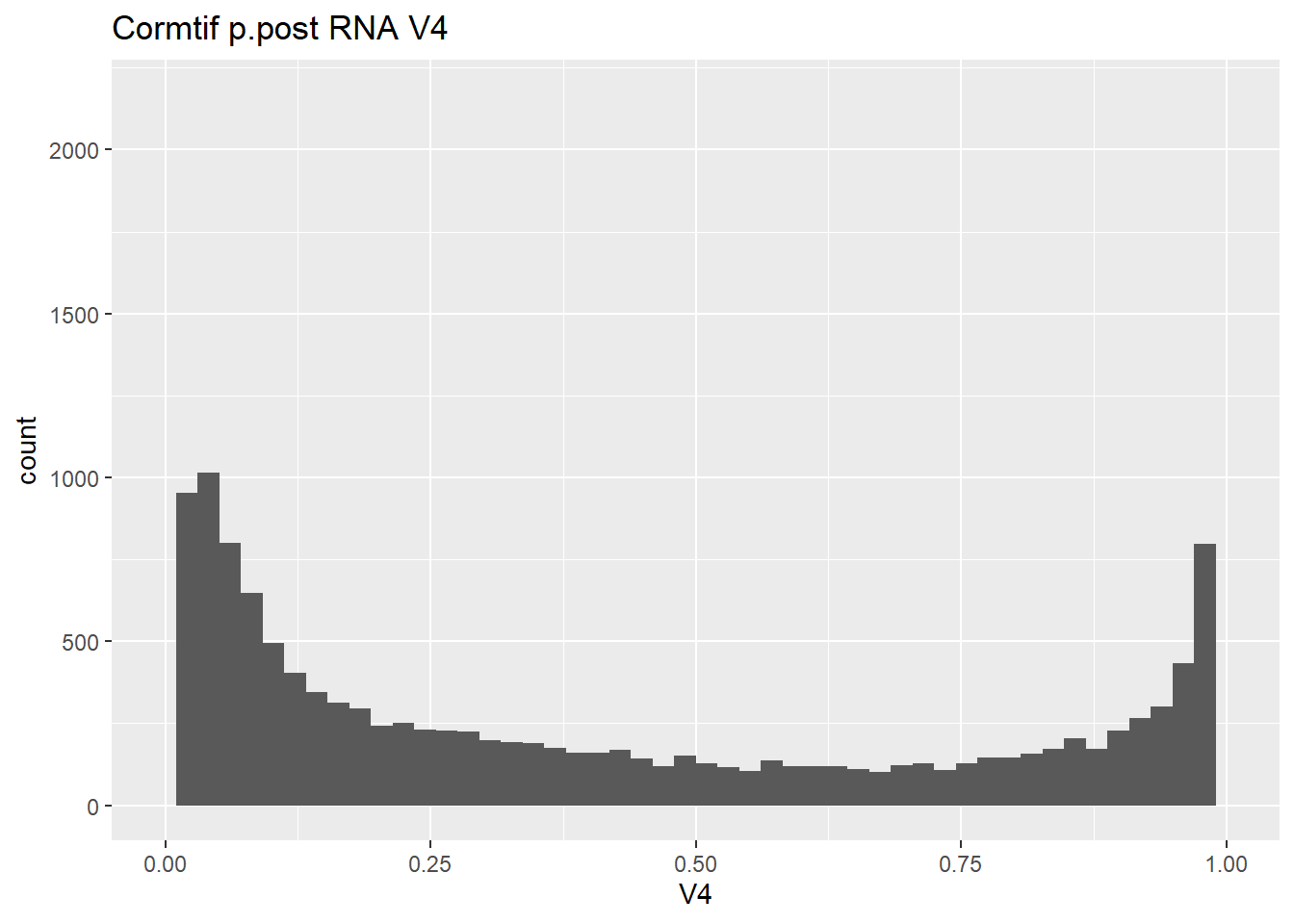
| Version | Author | Date |
|---|---|---|
| f388c90 | John D. Hurley | 2026-01-28 |
gene_prob_ppost %>%
as.data.frame() %>%
ggplot(., aes(x = V5))+
geom_histogram(bins = 50)+
xlim(0,1)+
ggtitle("Cormtif p.post RNA V5")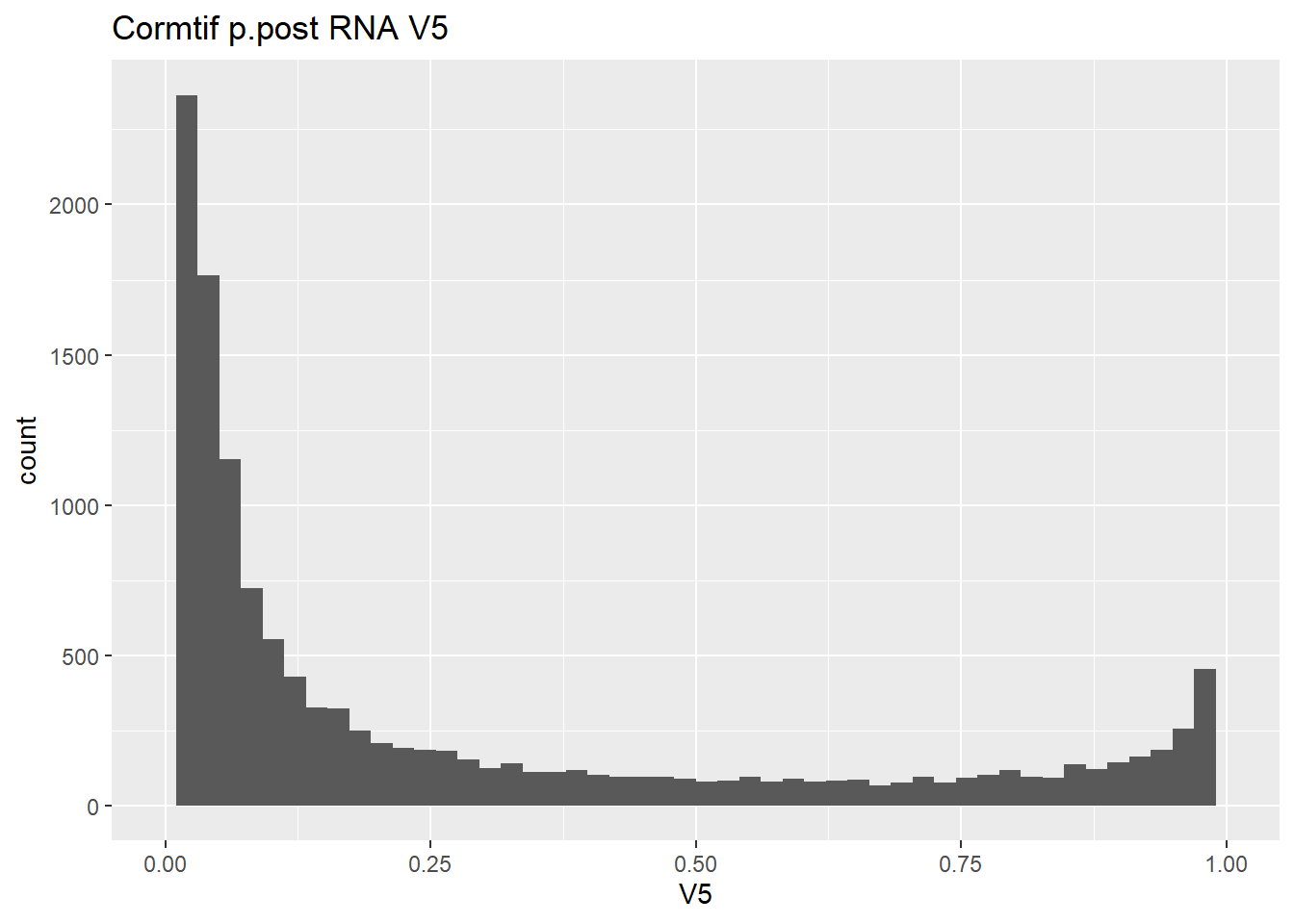
| Version | Author | Date |
|---|---|---|
| f388c90 | John D. Hurley | 2026-01-28 |
gene_prob_ppost %>%
as.data.frame() %>%
ggplot(., aes(x = V6))+
geom_histogram(bins = 50)+
xlim(0,1)+
ggtitle("Cormtif p.post RNA V6")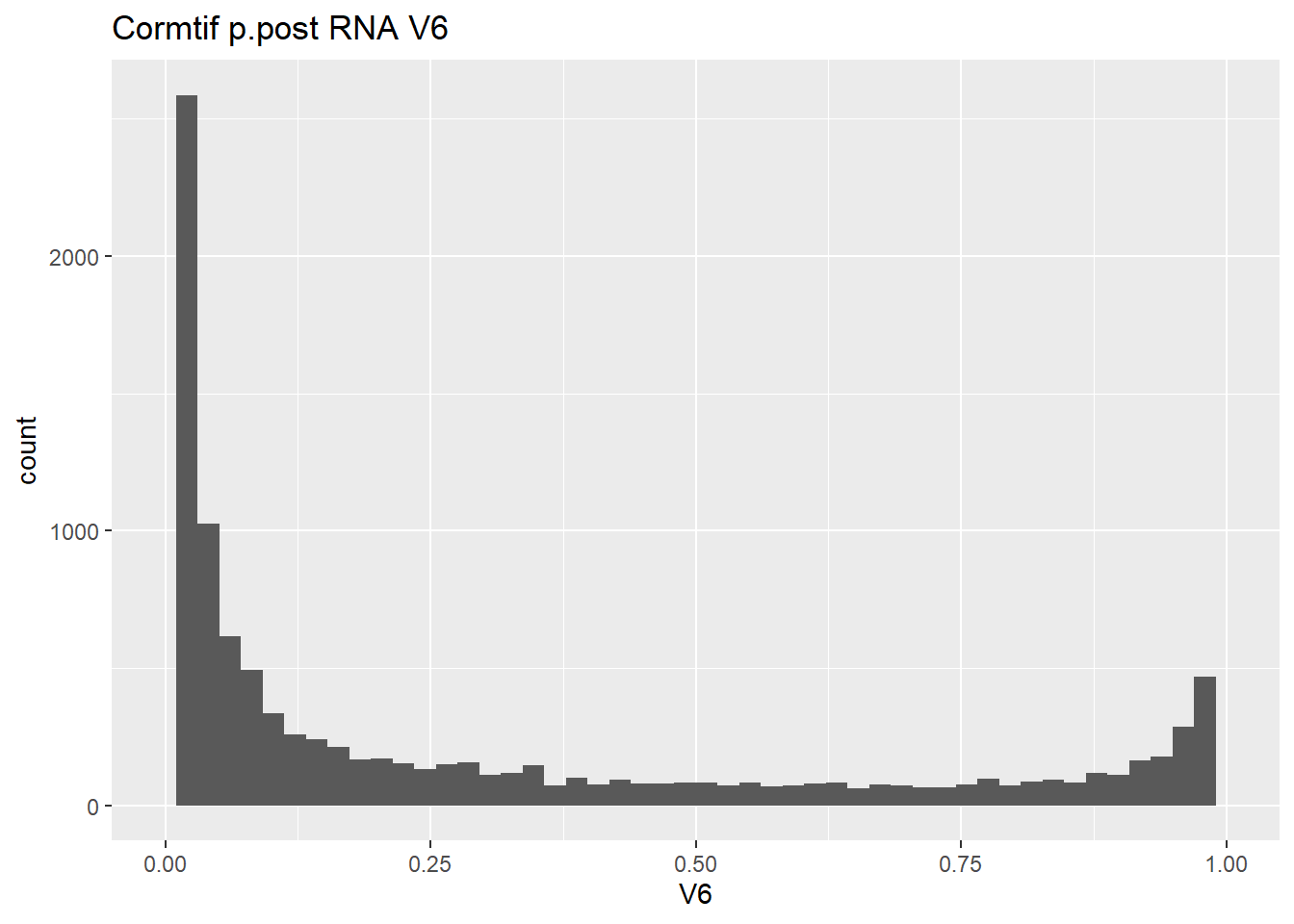
| Version | Author | Date |
|---|---|---|
| f388c90 | John D. Hurley | 2026-01-28 |
gene_prob_ppost %>%
as.data.frame() %>%
ggplot(., aes(x = V7))+
geom_histogram(bins = 50)+
xlim(0,1)+
ggtitle("Cormtif p.post RNA V7")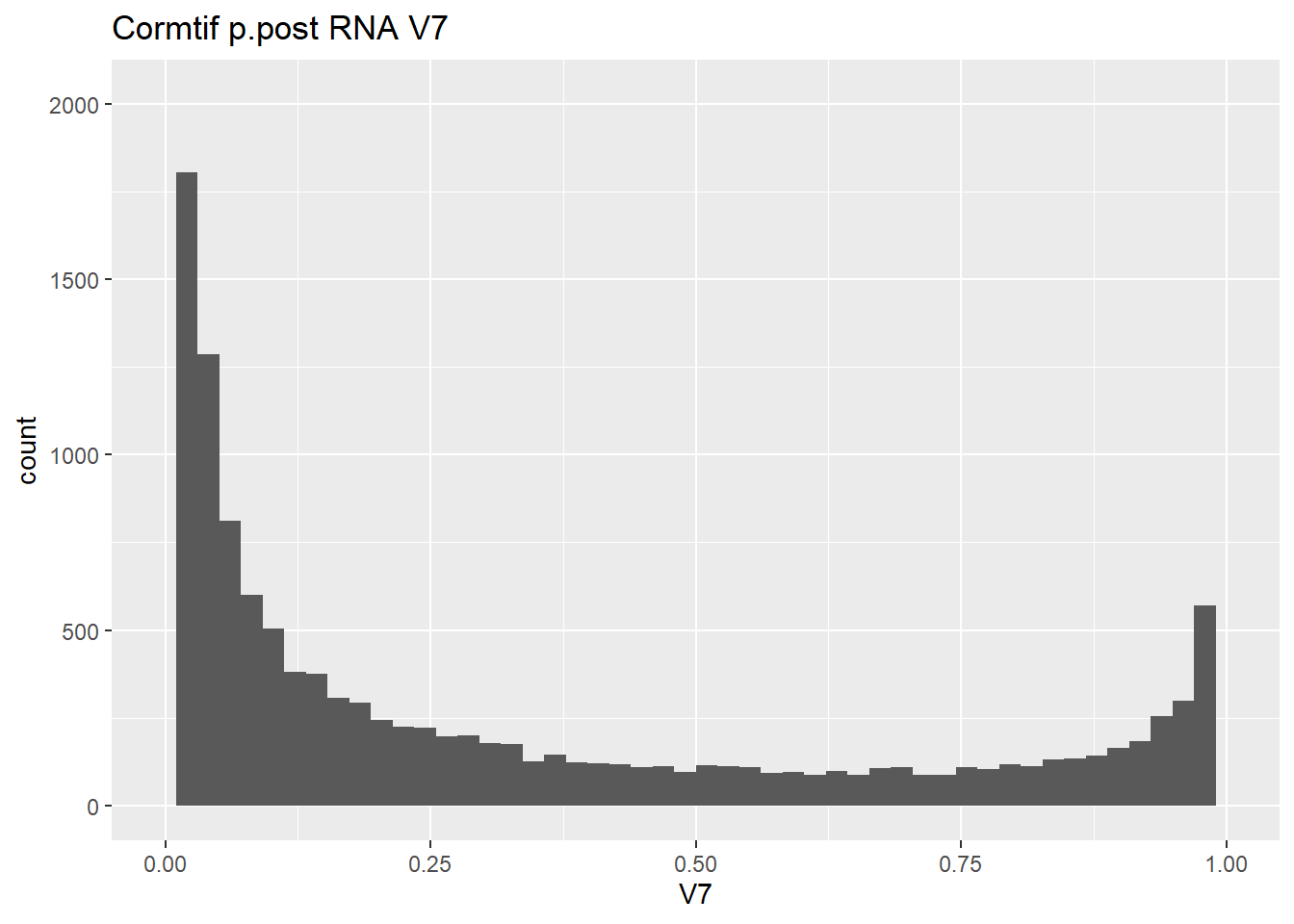
| Version | Author | Date |
|---|---|---|
| f388c90 | John D. Hurley | 2026-01-28 |
gene_prob_ppost %>%
as.data.frame() %>%
ggplot(., aes(x = V8))+
geom_histogram(bins = 50)+
xlim(0,1)+
ggtitle("Cormtif p.post RNA V8")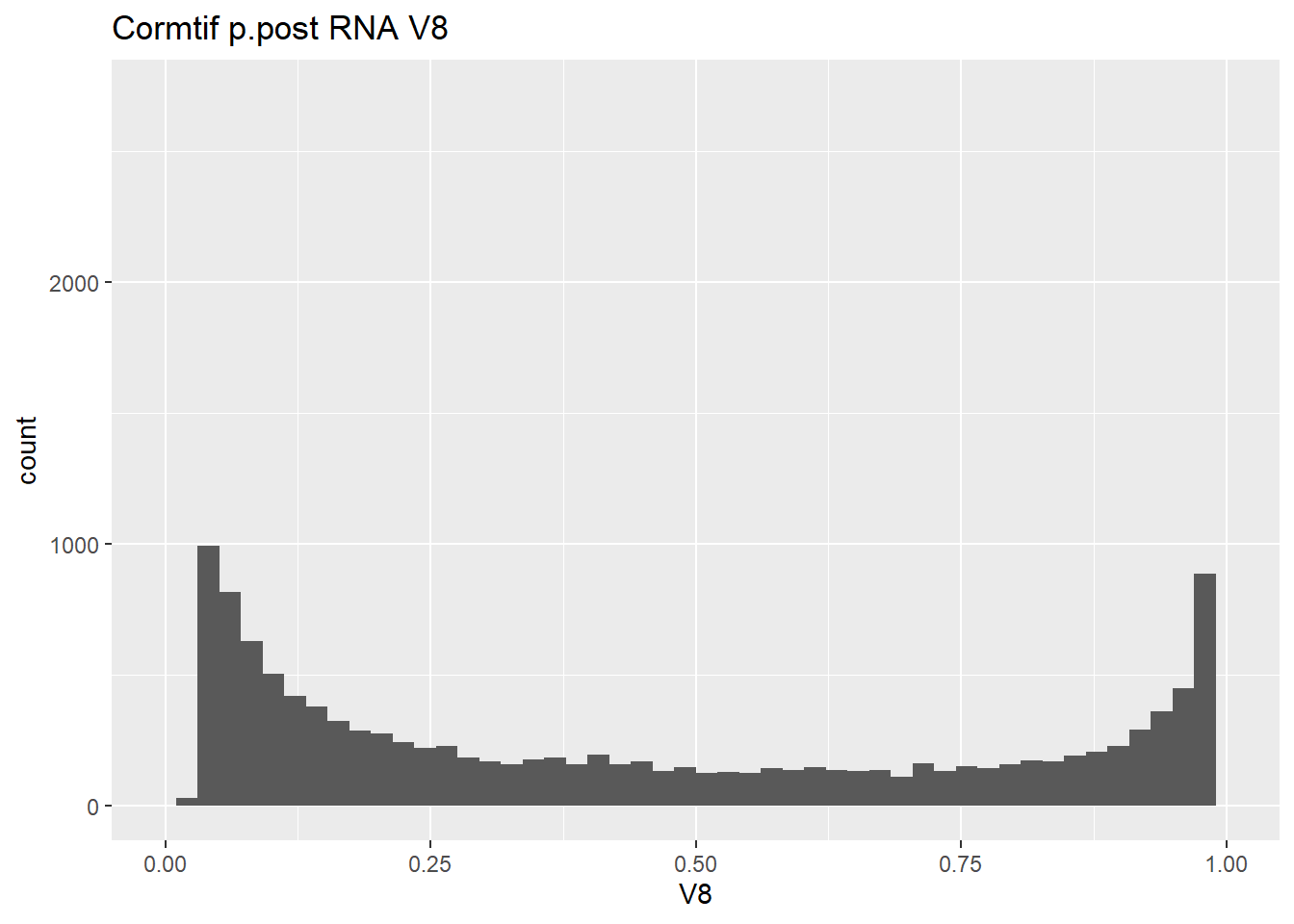
| Version | Author | Date |
|---|---|---|
| f388c90 | John D. Hurley | 2026-01-28 |
motif.fit <- motif.fitted
log2cpm <- Filt_RMG0_RNA_log2cpm_NoD4
#extract the cluster likelihood - which DEGs are most likely to be in this cluster
motif_prob <- motif.fit$bestmotif$clustlike
rownames(motif_prob) <- rownames(gene_prob_ppost)
# saveRDS(motif_prob,"data/RNA_motif_prob_Trajectory.RDS")
# motif.fitted$bestmotif$clustlike
clust1_RNA <-
motif_prob %>%
as.data.frame() %>%
filter(V1 > 0.5 & V2 < 0.5 & V3 < 0.5 & V4 < 0.5 & V5 < 0.5 & V6 < 0.5 & V7 < 0.5 & V8 < 0.5 & V9 < 0.5 & V10 < 0.5 & V11 < 0.5) %>%
rownames()
print("clust1_RNA")[1] "clust1_RNA"length(clust1_RNA)[1] 3381clust2_RNA <-
motif_prob %>%
as.data.frame() %>%
filter(V1 < 0.5 & V2 > 0.5 & V3 < 0.5 & V4 < 0.5 & V5 < 0.5 & V6 < 0.5 & V7 < 0.5 & V8 < 0.5 & V9 < 0.5 & V10 < 0.5 & V11 < 0.5) %>%
rownames()
print("clust2_RNA")[1] "clust2_RNA"length(clust2_RNA)[1] 815clust3_RNA <-
motif_prob %>%
as.data.frame() %>%
filter(V1 < 0.5 & V2 < 0.5 & V3 > 0.5 & V4 < 0.5 & V5 < 0.5 & V6 < 0.5 & V7 < 0.5 & V8 < 0.5 & V9 < 0.5 & V10 < 0.5 & V11 < 0.5) %>%
rownames()
print("clust3_RNA")[1] "clust3_RNA"length(clust3_RNA)[1] 623clust4_RNA <-
motif_prob %>%
as.data.frame() %>%
filter(V1 < 0.5 & V2 < 0.5 & V3 < 0.5 & V4 > 0.5 & V5 < 0.5 & V6 < 0.5 & V7 < 0.5 & V8 < 0.5 & V9 < 0.5 & V10 < 0.5) %>%
rownames()
print("clust4_RNA")[1] "clust4_RNA"length(clust4_RNA)[1] 1129clust5_RNA <-
motif_prob %>%
as.data.frame() %>%
filter(V1 < 0.5 & V2 < 0.5 & V3 < 0.5 & V4 < 0.5 & V5 > 0.5 & V6 < 0.5 & V7 < 0.5 & V8 < 0.5 & V9 < 0.5 & V10 < 0.5 & V11 < 0.5) %>%
rownames()
print("clust5_RNA")[1] "clust5_RNA"length(clust5_RNA)[1] 37clust6_RNA <-
motif_prob %>%
as.data.frame() %>%
filter(V1 < 0.5 & V2 < 0.5 & V3 < 0.5 & V4 < 0.5 & V5 < 0.5 & V6 > 0.5 & V7 < 0.5 & V8 < 0.5 & V9 < 0.5 & V10 < 0.5 & V11 < 0.5) %>%
rownames()
print("clust6_RNA")[1] "clust6_RNA"length(clust6_RNA)[1] 416clust7_RNA <-
motif_prob %>%
as.data.frame() %>%
filter(V1 < 0.5 & V2 < 0.5 & V3 < 0.5 & V4 < 0.5 & V5 < 0.5 & V6 < 0.5 & V7 > 0.5 & V8 < 0.5 & V9 < 0.5 & V10 < 0.5 & V11 < 0.5) %>%
rownames()
print("clust7_RNA")[1] "clust7_RNA"length(clust7_RNA)[1] 586clust8_RNA <-
motif_prob %>%
as.data.frame() %>%
filter(V1 < 0.5 & V2 < 0.5 & V3 < 0.5 & V4 < 0.5 & V5 <0.5 & V6 < 0.5 & V7 < 0.5 & V8 > 0.5 & V9 < 0.5 & V10 < 0.5 & V11 < 0.5) %>%
rownames()
print("clust8_RNA")[1] "clust8_RNA"length(clust8_RNA)[1] 628clust9_RNA <-
motif_prob %>%
as.data.frame() %>%
filter(V1 < 0.5 & V2 < 0.5 & V3 < 0.5 & V4 < 0.5 & V5 <0.5 & V6 < 0.5 & V7 < 0.5 & V8 < 0.5 & V9 > 0.5 & V10 < 0.5 & V11 < 0.5) %>%
rownames()
print("clust9_RNA")[1] "clust9_RNA"length(clust9_RNA)[1] 669clust10_RNA <-
motif_prob %>%
as.data.frame() %>%
filter(V1 < 0.5 & V2 < 0.5 & V3 < 0.5 & V4 < 0.5 & V5 <0.5 & V6 < 0.5 & V7 < 0.5 & V8 < 0.5 & V9 < 0.5 & V10 > 0.5 & V11 < 0.5) %>%
rownames()
print("clust10_RNA")[1] "clust10_RNA"length(clust10_RNA)[1] 1560clust11_RNA <-
motif_prob %>%
as.data.frame() %>%
filter(V1 < 0.5 & V2 < 0.5 & V3 < 0.5 & V4 < 0.5 & V5 <0.5 & V6 < 0.5 & V7 < 0.5 & V8 < 0.5 & V9 < 0.5 & V10 < 0.5 & V11 > 0.5) %>%
rownames()
print("clust11_RNA")[1] "clust11_RNA"length(clust11_RNA)[1] 656clust_names_list <- list(
clust1_RNA,
clust2_RNA,
clust3_RNA,
clust4_RNA,
clust5_RNA,
clust6_RNA,
clust7_RNA,
clust8_RNA,
clust9_RNA,
clust10_RNA,
clust11_RNA
)
names(clust_names_list) <- paste0("clust", 1:11)
print(overlap_matirx <- check_overlap(clust_names_list))$clust1
clust1 clust2 clust3 clust4 clust5 clust6 clust7 clust8 clust9 clust10
NA 0 0 0 0 0 0 0 0 0
clust11
0
$clust2
clust1 clust2 clust3 clust4 clust5 clust6 clust7 clust8 clust9 clust10
0 NA 0 0 0 0 0 0 0 0
clust11
0
$clust3
clust1 clust2 clust3 clust4 clust5 clust6 clust7 clust8 clust9 clust10
0 0 NA 0 0 0 0 0 0 0
clust11
0
$clust4
clust1 clust2 clust3 clust4 clust5 clust6 clust7 clust8 clust9 clust10
0 0 0 NA 0 0 0 0 0 0
clust11
0
$clust5
clust1 clust2 clust3 clust4 clust5 clust6 clust7 clust8 clust9 clust10
0 0 0 0 NA 0 0 0 0 0
clust11
0
$clust6
clust1 clust2 clust3 clust4 clust5 clust6 clust7 clust8 clust9 clust10
0 0 0 0 0 NA 0 0 0 0
clust11
0
$clust7
clust1 clust2 clust3 clust4 clust5 clust6 clust7 clust8 clust9 clust10
0 0 0 0 0 0 NA 0 0 0
clust11
0
$clust8
clust1 clust2 clust3 clust4 clust5 clust6 clust7 clust8 clust9 clust10
0 0 0 0 0 0 0 NA 0 0
clust11
0
$clust9
clust1 clust2 clust3 clust4 clust5 clust6 clust7 clust8 clust9 clust10
0 0 0 0 0 0 0 0 NA 0
clust11
0
$clust10
clust1 clust2 clust3 clust4 clust5 clust6 clust7 clust8 clust9 clust10
0 0 0 0 0 0 0 0 0 NA
clust11
0
$clust11
clust1 clust2 clust3 clust4 clust5 clust6 clust7 clust8 clust9 clust10
0 0 0 0 0 0 0 0 0 0
clust11
NA #View(motif_prob)
# write.csv(clust1_RNA,"data/Cormotif/clust1_RNA_Trajectory.csv")
# write.csv(clust2_RNA,"data/Cormotif/clust2_RNA_Trajectory.csv")
# write.csv(clust3_RNA,"data/Cormotif/clust3_RNA_Trajectory.csv")
# write.csv(clust4_RNA,"data/Cormotif/clust4_RNA_Trajectory.csv")
# write.csv(clust5_RNA,"data/Cormotif/clust5_RNA_Trajectory.csv")
# write.csv(clust6_RNA,"data/Cormotif/clust6_RNA_Trajectory.csv")
# write.csv(clust7_RNA,"data/Cormotif/clust7_RNA_Trajectory.csv")
# write.csv(clust8_RNA,"data/Cormotif/clust8_RNA_Trajectory.csv")
# write.csv(clust9_RNA,"data/Cormotif/clust9_RNA_Trajectory.csv")
# write.csv(clust10_RNA,"data/Cormotif/clust10_RNA_Trajectory.csv")
# write.csv(clust11_RNA,"data/Cormotif/clust11_RNA_Trajectory.csv")
# saveRDS(clust1_RNA,"data/Cormotif/clust1_RNA_Trajectory.RDS")
# saveRDS(clust2_RNA,"data/Cormotif/clust2_RNA_Trajectory.RDS")
# saveRDS(clust3_RNA,"data/Cormotif/clust3_RNA_Trajectory.RDS")
# saveRDS(clust4_RNA,"data/Cormotif/clust4_RNA_Trajectory.RDS")
# saveRDS(clust5_RNA,"data/Cormotif/clust5_RNA_Trajectory.RDS")
# saveRDS(clust6_RNA,"data/Cormotif/clust6_RNA_Trajectory.RDS")
# saveRDS(clust7_RNA,"data/Cormotif/clust7_RNA_Trajectory.RDS")
# saveRDS(clust8_RNA,"data/Cormotif/clust8_RNA_Trajectory.RDS")
# saveRDS(clust9_RNA,"data/Cormotif/clust9_RNA_Trajectory.RDS")
# saveRDS(clust10_RNA,"data/Cormotif/clust10_RNA_Trajectory.RDS")
# saveRDS(clust1_RNA,"data/Cormotif/clust11_RNA_Trajectory.RDS")
# ```# git -> commit all changes
# git -> push
# wflow_publish("analysis/RNA_CorMotif_Trajectory_Ensemble.Rmd")
sessionInfo()R version 4.5.1 (2025-06-13 ucrt)
Platform: x86_64-w64-mingw32/x64
Running under: Windows 11 x64 (build 26100)
Matrix products: default
LAPACK version 3.12.1
locale:
[1] LC_COLLATE=English_United States.utf8
[2] LC_CTYPE=English_United States.utf8
[3] LC_MONETARY=English_United States.utf8
[4] LC_NUMERIC=C
[5] LC_TIME=English_United States.utf8
time zone: America/Chicago
tzcode source: internal
attached base packages:
[1] grid stats4 stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] ComplexHeatmap_2.24.1 ggfortify_0.4.19
[3] readxl_1.4.5 RUVSeq_1.42.0
[5] EDASeq_2.42.0 ShortRead_1.66.0
[7] GenomicAlignments_1.44.0 SummarizedExperiment_1.38.1
[9] MatrixGenerics_1.20.0 matrixStats_1.5.0
[11] Rsamtools_2.24.0 GenomicRanges_1.60.0
[13] Biostrings_2.76.0 GenomeInfoDb_1.44.3
[15] XVector_0.48.0 BiocParallel_1.42.1
[17] lubridate_1.9.4 forcats_1.0.1
[19] stringr_1.5.2 purrr_1.1.0
[21] tidyr_1.3.1 tidyverse_2.0.0
[23] pheatmap_1.0.13 org.Hs.eg.db_3.21.0
[25] AnnotationDbi_1.70.0 IRanges_2.42.0
[27] S4Vectors_0.46.0 Biobase_2.68.0
[29] BiocGenerics_0.54.0 generics_0.1.4
[31] readr_2.1.5 ggrepel_0.9.6
[33] dplyr_1.1.4 tibble_3.3.0
[35] ggplot2_4.0.0 edgeR_4.6.3
[37] limma_3.64.3 workflowr_1.7.2
loaded via a namespace (and not attached):
[1] RColorBrewer_1.1-3 shape_1.4.6.1 rstudioapi_0.18.0
[4] jsonlite_2.0.0 magrittr_2.0.3 GenomicFeatures_1.60.0
[7] farver_2.1.2 rmarkdown_2.30 GlobalOptions_0.1.3
[10] BiocIO_1.18.0 fs_1.6.6 vctrs_0.6.5
[13] memoise_2.0.1 RCurl_1.98-1.17 htmltools_0.5.8.1
[16] S4Arrays_1.8.1 progress_1.2.3 curl_7.0.0
[19] cellranger_1.1.0 SparseArray_1.8.1 sass_0.4.10
[22] bslib_0.9.0 httr2_1.2.2 cachem_1.1.0
[25] whisker_0.4.1 iterators_1.0.14 lifecycle_1.0.5
[28] pkgconfig_2.0.3 Matrix_1.7-3 R6_2.6.1
[31] fastmap_1.2.0 clue_0.3-66 GenomeInfoDbData_1.2.14
[34] digest_0.6.37 colorspace_2.1-2 ps_1.9.1
[37] rprojroot_2.1.1 RSQLite_2.4.3 hwriter_1.3.2.1
[40] labeling_0.4.3 filelock_1.0.3 timechange_0.3.0
[43] httr_1.4.7 abind_1.4-8 compiler_4.5.1
[46] doParallel_1.0.17 bit64_4.6.0-1 withr_3.0.2
[49] S7_0.2.0 DBI_1.2.3 R.utils_2.13.0
[52] biomaRt_2.64.0 MASS_7.3-65 rappdirs_0.3.4
[55] DelayedArray_0.34.1 rjson_0.2.23 tools_4.5.1
[58] otel_0.2.0 httpuv_1.6.16 R.oo_1.27.1
[61] glue_1.8.0 restfulr_0.0.16 callr_3.7.6
[64] promises_1.3.3 getPass_0.2-4 cluster_2.1.8.1
[67] gtable_0.3.6 tzdb_0.5.0 R.methodsS3_1.8.2
[70] hms_1.1.4 xml2_1.5.1 foreach_1.5.2
[73] pillar_1.11.1 later_1.4.4 circlize_0.4.17
[76] BiocFileCache_2.16.2 lattice_0.22-7 rtracklayer_1.68.0
[79] aroma.light_3.38.0 bit_4.6.0 deldir_2.0-4
[82] tidyselect_1.2.1 locfit_1.5-9.12 knitr_1.51
[85] git2r_0.36.2 gridExtra_2.3 xfun_0.53
[88] statmod_1.5.0 stringi_1.8.7 UCSC.utils_1.4.0
[91] yaml_2.3.10 evaluate_1.0.5 codetools_0.2-20
[94] interp_1.1-6 cli_3.6.5 processx_3.8.6
[97] jquerylib_0.1.4 Rcpp_1.1.0 dbplyr_2.5.1
[100] png_0.1-8 XML_3.99-0.20 parallel_4.5.1
[103] blob_1.3.0 prettyunits_1.2.0 latticeExtra_0.6-31
[106] jpeg_0.1-11 bitops_1.0-9 pwalign_1.4.0
[109] scales_1.4.0 crayon_1.5.3 GetoptLong_1.1.0
[112] rlang_1.1.6 KEGGREST_1.48.1